Intelligence Archive
"How are mitochondrial genomes spatially organized within the mitochondrial network and faithfully inherited during cell division?"
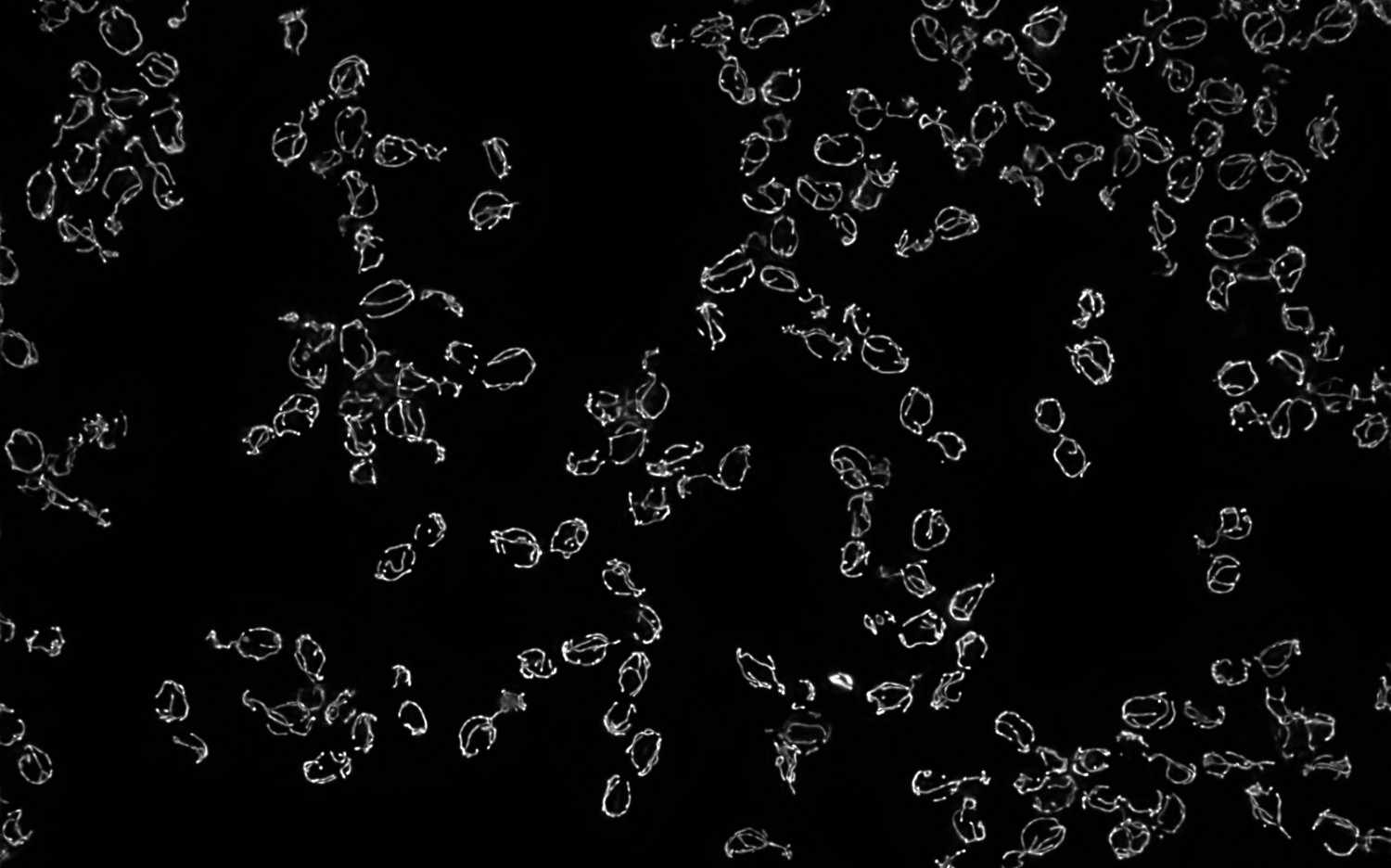
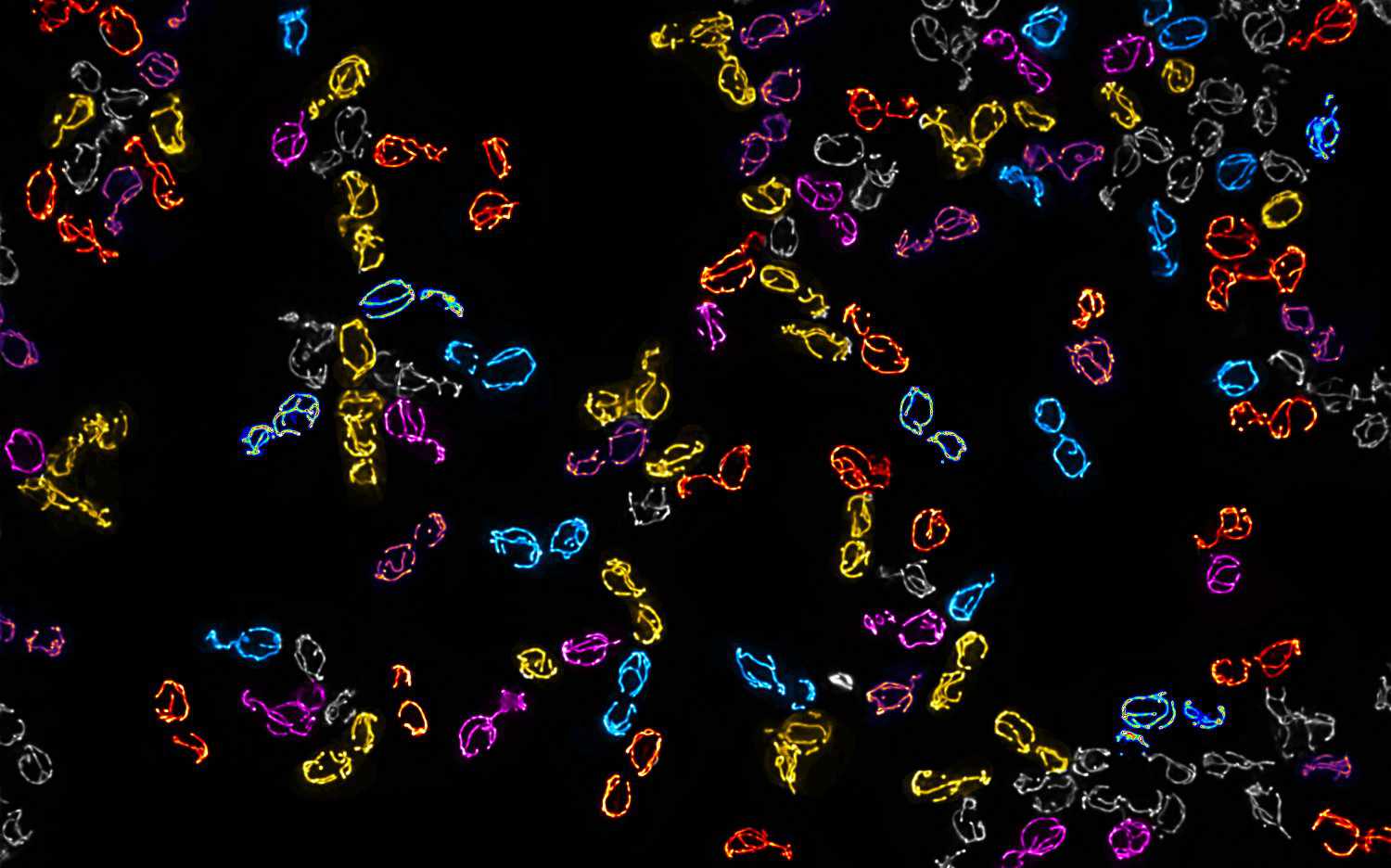
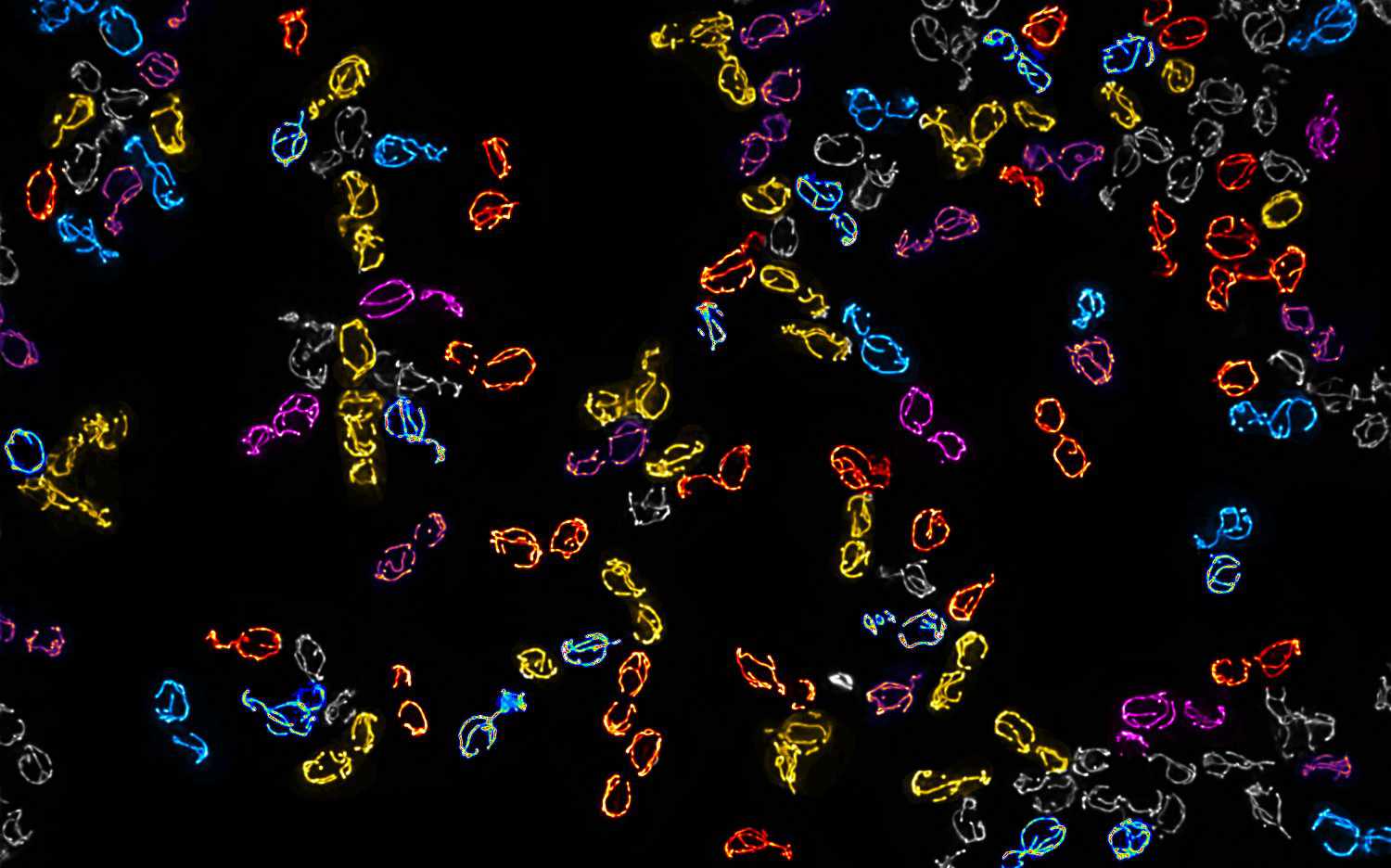
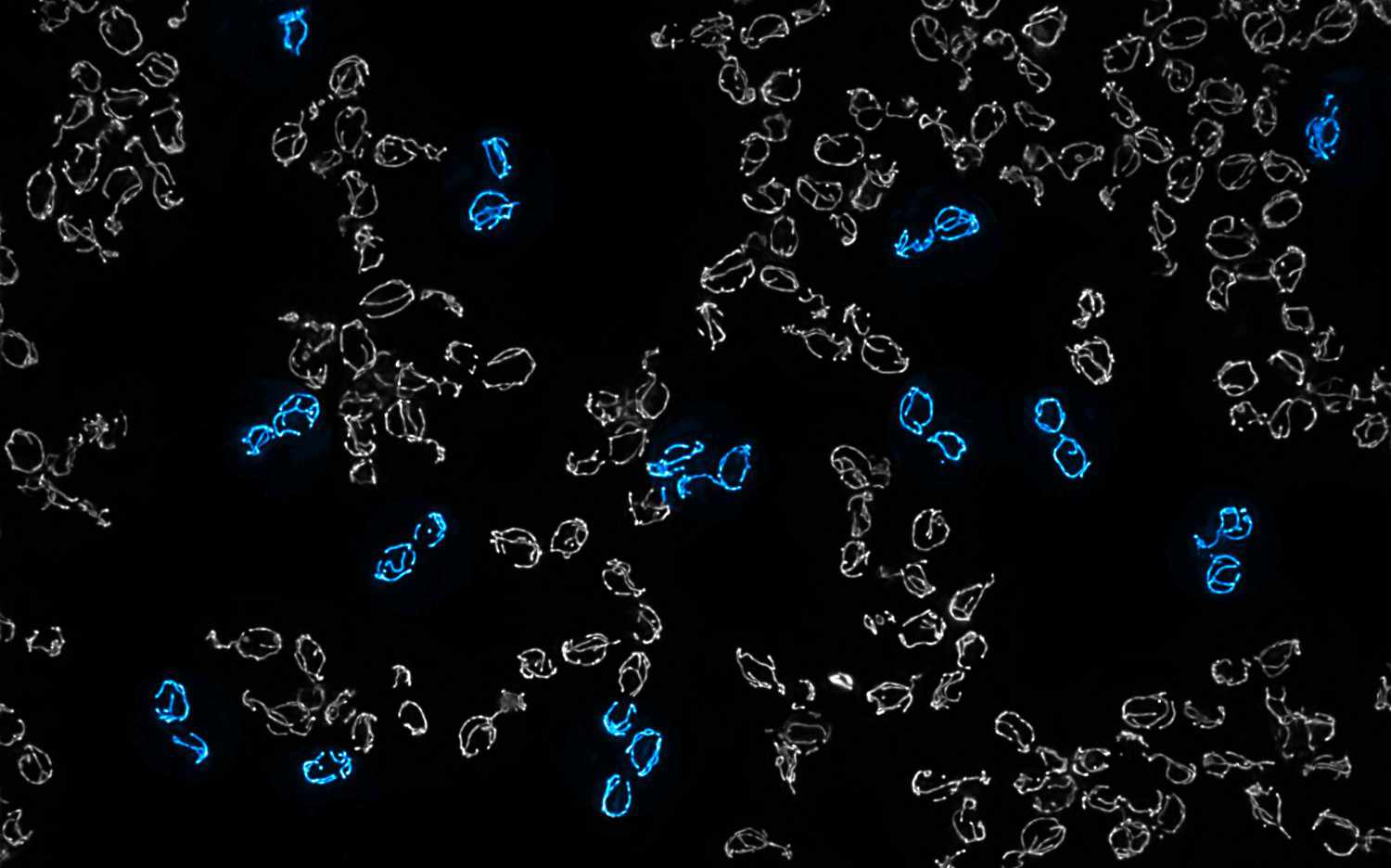
Mitochondrial DNA molecules are packaged into nucleoprotein structures called nucleoids and distributed throughout the dynamic mitochondrial network. The spatial organization of these nucleoids influences genome …
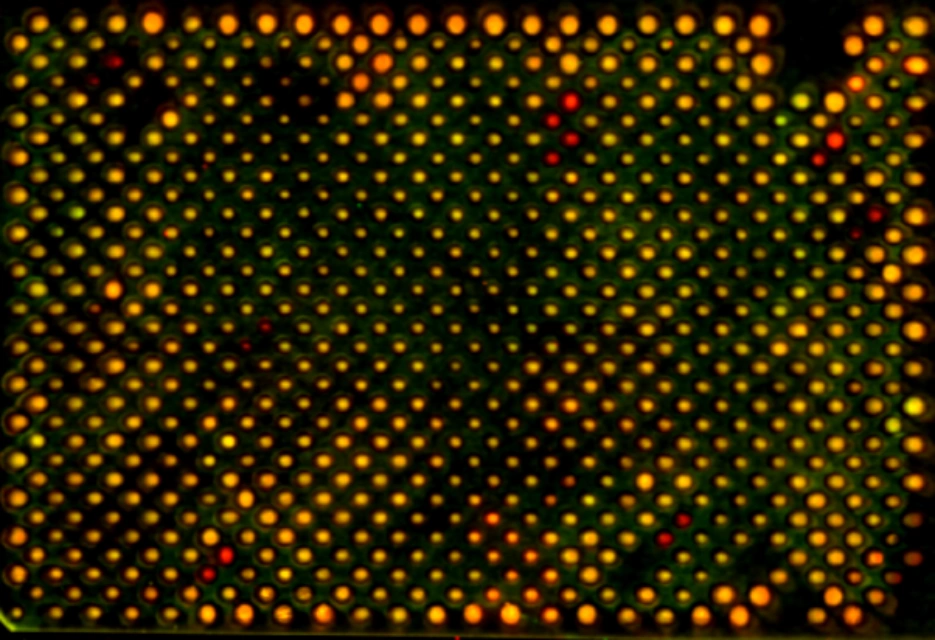

"How is the number of mitochondrial DNA molecules controlled and coordinated with cellular growth and metabolic demand?"
Each cell contains multiple copies of mitochondrial DNA, and maintaining the correct number of genomes is essential for proper mitochondrial function. Too few copies can …
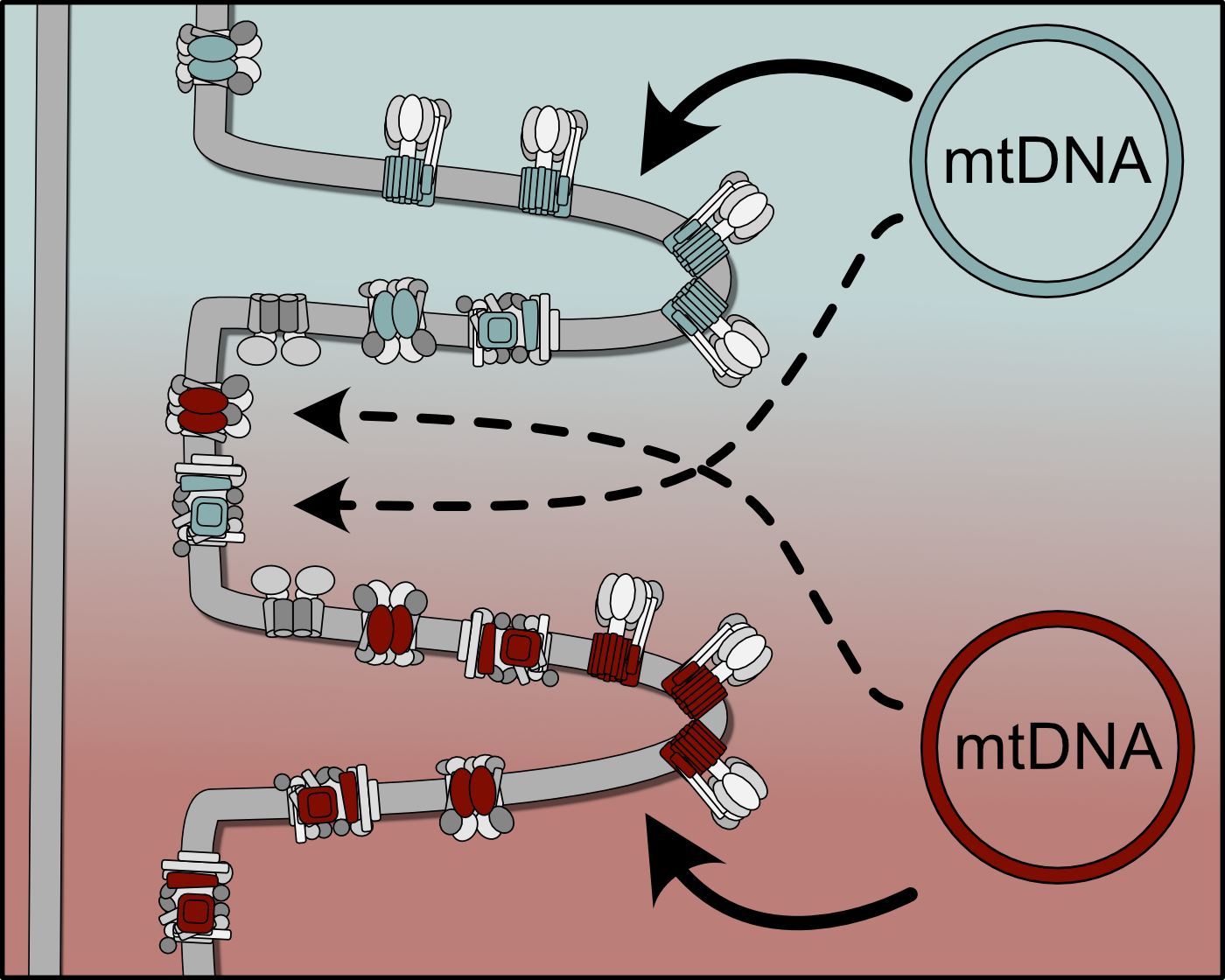
"How do cells detect and selectively eliminate defective mitochondrial genomes while preserving functional mtDNA?"
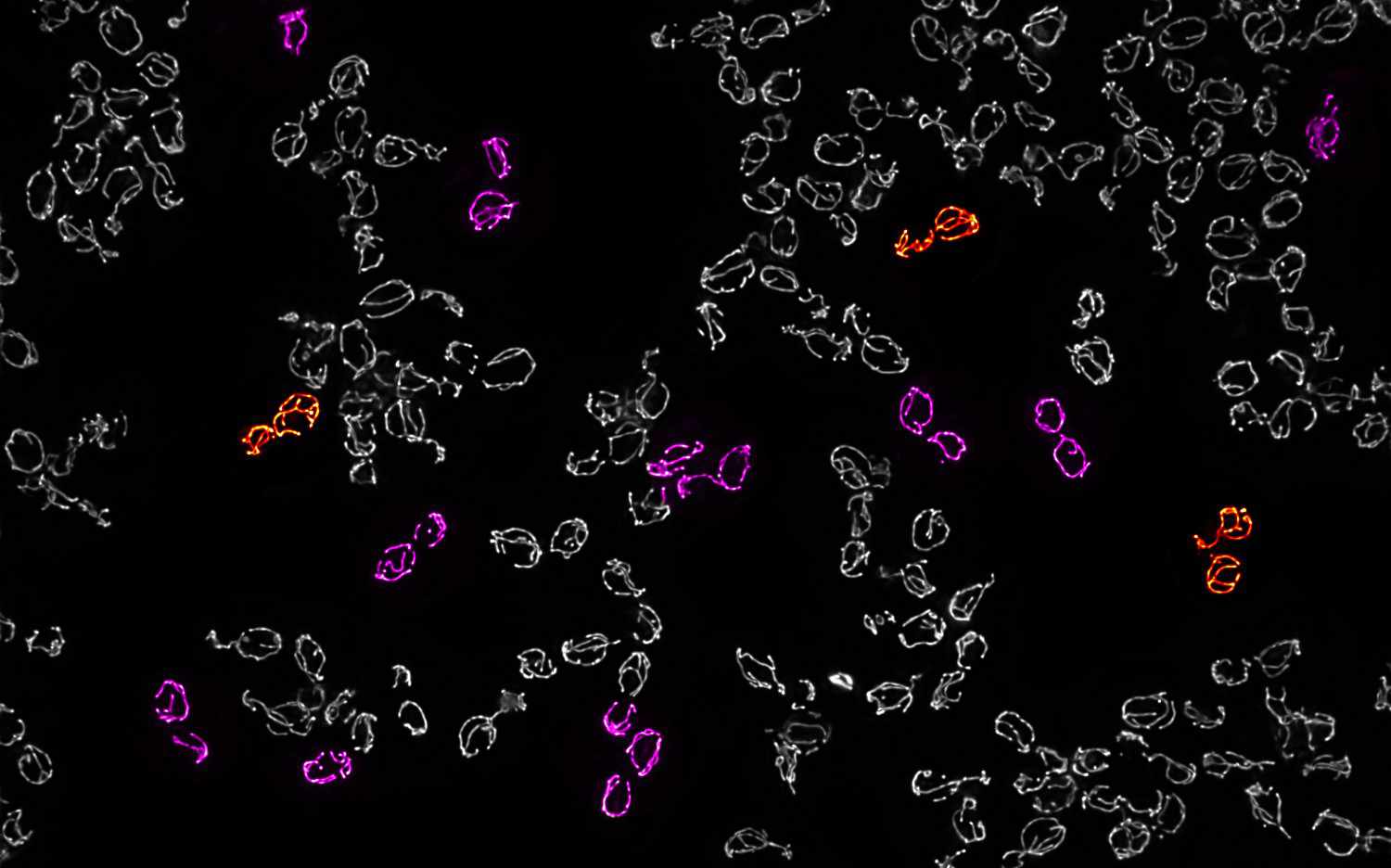
Mitochondrial DNA (mtDNA) is continuously exposed to damage and mutations due to its proximity to reactive metabolic processes. The accumulation of dysfunctional mtDNA molecules can …
The Osman Lab investigates mitochondrial genome maintenance in S. cerevisiae.
Audio summaries are generated by NotebookLM. Please consult the primary literature for final accuracy.
Mrx6 binds the Lon protease Pim1 N-terminal domain to confer selective substrate specificity and regulate mtDNA copy number
Schrott, S., Marafelli, I., Gerle, C., Bibinger, S., Bertgen, L., Marino, G., Schwenkert, S., Rathberger, R., Herrmann, J., Osman, C.#
Local mitochondrial physiology defined by mtDNA quality guides purifying selection
Thoma, F., Hagen, J., Rathberger, R., Padovani, F., Hörl, D., Schmoller, K., Osman, C.#
Real-time assessment of mitochondrial DNA heteroplasmy dynamics at the single-cell level
Roussou, R., Metzler, D., Padovani, F., Thoma, F., Schwarz, R., Shraiman, B., Schmoller, K., Osman, C.#
Schrott, S., Osman, C.#
Cristae-dependent quality control of the mitochondrial genome
Jakubke, C.*, Roussou, R.*, Maiser, A., Schug, C., Thoma, F., Bunk, R., Hörl, D., Leonhardt, H., Walter, P., Klecker, T., Osman, C.#
Mrx6 regulates mitochondrial DNA copy number in Saccharomyces cerevisiae by engaging the evolutionarily conserved Lon protease Pim1
Göke, A., Schrott, S., Mizrak, A., Belyy, V., Osman, C.#, Walter, P.#
Integrity of the yeast mitochondrial genome, but not its distribution and inheritance, relies on mitochondrial fission and fusion
Osman, C., Noriega, T., Okreglak, V., Walter, P.
We are always enthusiastic about new collaborations and scientific exchange. While we may not always have funded vacancies, we are always interested in prospective postdocs and fellows who wish to apply for their own external funding. We are happy to discuss the possibility of hosting and supporting your fellowship application.
